Home

With the resources of the SUNY Research Foundation, and our history of successful partnerships, we are here to help move biomedical products and ideas to market.

Our scientists and core facilities can help move discoveries into practice and technologies into the marketplace.

Upstate is home to top research facilities with highly specialized equipment and advanced instrumentation, to support research and product development.

We are here to create the relationships and partnerships needed to move innovative ideas forward.
Upstate Biotech Ventures
In a partnership between Empire State Development, Upstate Medical University, the SUNY Research Foundation, and Excell Partners, the newly-launched Upstate Biotech Ventures invests in high-potential startups and small businesses affiliated with Upstate Medical University to drive research and technology innovation.
Recent Tech from SUNY Upstate
This technology uses specially designed sulfonium lipid nanoparticles to deliver mRNA directly to lu...
This technology uses specially designed sulfonium lipid nanoparticles to deliver mRNA directly to lung cells through the nose, offering a non-invasive, more effective and targeted treatment for pulmonary diseases like asthma and cystic fibrosis. Background:
The field of pulmonary medicine faces significant challenges in the treatment of diseases such as cystic fibrosis, asthma, and acute respiratory distress syndrome, all of which involve complex interactions between lung epithelial and immune cells. Advances in genetic medicine, particularly messenger RNA (mRNA) therapeutics, have opened new avenues for treating these conditions by enabling the direct modulation of gene expression within target cells. However, the success of mRNA-based therapies hinges on the ability to deliver these fragile molecules efficiently and specifically to the relevant lung cells. Intranasal administration is a promising route due to its non-invasiveness and direct access to the respiratory tract, but it requires delivery systems that can protect mRNA from degradation and ensure its uptake by the desired cell types. Current approaches to mRNA delivery, such as conventional lipid nanoparticles, face several limitations when applied to pulmonary diseases. These formulations often lack the specificity needed to target lung epithelial and immune cells, resulting in suboptimal therapeutic outcomes and potential off-target effects. Furthermore, many existing nanoparticles struggle to traverse the mucus barrier and are rapidly cleared from the respiratory tract, reducing the amount of mRNA that reaches the intended cells. Inefficient encapsulation and delivery can also lead to degradation of the mRNA payload before it exerts its therapeutic effect. As a result, there is a pressing need for more effective and targeted delivery vehicles that can overcome these biological barriers and improve the efficacy of mRNA-based treatments for lung diseases.Technology Overview:
This technology introduces a new class of sulfonium lipid nanoparticles specifically engineered for the intranasal delivery of mRNA molecules to lung epithelial and immune cells. These nanoparticles are carefully designed and synthesized to encapsulate and protect mRNA, ensuring efficient transport and localized release within the lungs. By optimizing the formulation, the technology achieves superior performance compared to existing lipid-based delivery systems, providing enhanced targeting and uptake by the intended lung cell populations. The system is designed to address the longstanding challenge of delivering genetic material directly to the respiratory tract, which is crucial for treating pulmonary diseases such as cystic fibrosis, asthma, and acute respiratory distress syndrome. What differentiates this technology is its unique chemical structure and formulation, which enable highly specific and efficient delivery of mRNA to lung cells via the intranasal route. Unlike conventional lipid nanoparticles, the sulfonium-based design offers improved stability, cellular uptake, and targeting capabilities, resulting in higher therapeutic efficacy and reduced off-target effects. The technology fills a critical gap in the market by providing a non-invasive, localized delivery method that can be readily adapted for a variety of mRNA-based therapeutics. Its development, supported by NIH funding and validated through rigorous experimentation and peer-reviewed disclosure, positions it as a leading solution for advancing pulmonary drug delivery and gene therapy. https://suny.technologypublisher.com/files/sites/adobestock_1237485827.jpegAdvantages:
• Enables targeted and efficient intranasal delivery of mRNA to lung epithelial and immune cells
• Optimized sulfonium lipid nanoparticles outperform existing benchmark lipid formulations for mRNA delivery
• Facilitates localized gene expression critical for treating pulmonary diseases such as cystic fibrosis, asthma, and acute respiratory distress syndrome
• Non-invasive administration route through intranasal delivery improves patient compliance
• Supports development of novel mRNA-based therapeutics for a wide range of lung diseases
• Innovative chemical design avoids reliance on existing intellectual property, allowing for broad application and further development Applications:
• mRNA therapeutics for cystic fibrosis
• Asthma gene therapy delivery
• Acute respiratory distress treatment
• Targeted lung cancer mRNA therapy
• Vaccines for respiratory infections Intellectual Property Summary:
Patent application: 63/783,376, filed on 4/4/2025
Issued patent
Know-how based
CopyrightStage of Development:
TRL 3
Sulfonium lipid nanoparticles are specifically engineered for the intranasal delivery of mRNA molecules to lung epithelial and immune cells Licensing Status:
This technology is available for licensing.
This technology uses fenretinide, a synthetic vitamin A derivative, to boost the body’s adaptive imm...
This technology uses fenretinide, a synthetic vitamin A derivative, to boost the body’s adaptive immune response after vaccination or infection, making vaccines more effective, especially for poorly immunogenic agents or immunocompromised individuals, without increasing infection severity. Background:
The field of vaccine development and immunotherapy is critical for controlling infectious diseases and protecting vulnerable populations. Effective vaccines rely on the body’s ability to mount a strong and lasting adaptive immune response, which is often facilitated by adjuvants—substances added to vaccines to enhance immunogenicity. However, many vaccines, particularly those targeting emerging pathogens or used in immunocompromised individuals, elicit suboptimal immune responses. This challenge is compounded in settings where dose sparing is necessary, such as during pandemics or when manufacturing capacity is limited. There is a growing need for novel strategies that can reliably boost immune responses, especially for populations with weakened immunity due to age, underlying disease, or immunosuppressive treatments. Current approaches to enhancing vaccine efficacy and immune responses face several limitations. Traditional adjuvants, such as aluminum salts or oil-in-water emulsions, can be associated with increased reactogenicity, complex manufacturing requirements, and regulatory hurdles, which slow down the development and deployment of new vaccines. Moreover, these adjuvants are not always effective for all vaccine types or in all patient populations, leaving gaps in protection for those most at risk. In immunocompromised individuals, even potent adjuvants may fail to elicit adequate immunity, and there are few safe, broadly applicable alternatives. As a result, there is a pressing need for new immune-enhancing approaches that are safe, effective across diverse populations, and compatible with existing vaccine platforms.Technology Overview:
This technology utilizes fenretinide (N-(4-hydroxyphenyl)retinamide), a synthetic derivative of vitamin A, as a universal immune adjuvant to enhance adaptive immune responses following vaccination or pathogen exposure. When administered after inoculation, fenretinide specifically increases serum levels of key cytokines—IP-10 and IFN-gamma—both of which are critical for T cell activation and effector function. Importantly, this immunomodulatory effect occurs without altering the course of infection or affecting viral titers, as demonstrated in clinical trials involving dengue virus. The approach is particularly valuable in scenarios where vaccines are poorly immunogenic, where individuals are immunocompromised, or where dose sparing is necessary for safety or production efficiency. Fenretinide’s favorable toxicity profile, established through its prior use in cancer prevention, further supports its suitability for broad application as an immune response enhancer. What differentiates this technology is its unique mechanism of action and versatility. Unlike traditional adjuvants, which often require complex development and can introduce regulatory and manufacturing challenges, fenretinide acts post-inoculation to selectively boost adaptive immunity without increasing inflammation or viral replication. This allows for improved immune responses even in vulnerable populations, such as the elderly or immunosuppressed, and enables more efficient use of vaccine doses. Additionally, its ability to function as an adjuvant substitute offers significant advantages in terms of reducing research and development costs and streamlining regulatory approval. The method’s demonstrated efficacy in human challenge models, combined with its broad applicability and established safety, positions it as a transformative solution for enhancing vaccine performance and infectious disease management. https://suny.technologypublisher.com/files/sites/adobestock_291684715.jpeg
Photo for reference only, not a depiction of the invention.Advantages:
• Enhances adaptive immune responses by increasing key cytokines (IP-10 and IFN-gamma) associated with T cell activation and effector function
• Acts as a universal immune adjuvant effective when administered post-inoculation with pathogens or vaccines
• Improves immune responses to poorly immunogenic vaccines and immunostimulatory agents
• Boosts immunity in immunocompromised individuals, including those undergoing chemotherapy or radiation
• Enables dose sparing, reducing the amount of vaccine or immunostimulant needed
• Provides an alternative to traditional adjuvants, potentially lowering R&D, manufacturing, and regulatory costs
• Does not alter the course of infection or viral titers, focusing on immune enhancement rather than direct antiviral effects Applications:
• Universal vaccine immune adjuvant
• Enhancing immunity in immunocompromised
• Dose sparing for vaccines
• Adjuvant substitute for vaccine development Intellectual Property Summary:
Patent application 63/861,507 filed 8/11/2025Stage of Development:
TRL 7
The technology is at an early clinical development stage, supported by human challenge study data demonstrating enhanced adaptive immune responses (increased IP 10 and IFN γ) without altering infection course or viral titers. Further clinical evaluation in targeted vaccine and infectious disease indications would advance it toward TRL 8.Licensing Status:
This technology is available for licensing.
A surgical "sewing machine" for rapid graft quilting and suturing in challenging spaces. Background:...
A surgical "sewing machine" for rapid graft quilting and suturing in challenging spaces. Background: Grafts are commonly employed in urologic reconstructive surgery, but anchoring them in less accessible areas -- as in luminal stenosis surgery -- can be difficult. A novel surgical "sewing machine" capable of quilting and suturing in tight spaces was developed to help solve this problem. Technology Overview: The repair of luminal stenosis involves an incision through the stenosed segments and followed by the application of a buccal mucosa graft to serve as protection. A surgical sewing machine was assembled by threading absorbable 4-0 barbed suture through a 20-gauge hollow needle. The result is rapid, one hand suturing for graft quilting, with a running stitch, akin to the way a sewing machine makes a continuous stitch across the hem of a skirt. This suturing device, developed by Upstate Medical University researchers, has been used in posterior urethroplasties, a transvesical bladder neck reconstruction, augmented perineal urethrostomy, and a vaginoplasty revision. In each case, the graft survived and there was no recurrence of disease. Advantages:
- Higher rate of graft success with no recurrence of disease
- One handed suturing avoids alternating movements to reposition the needle, and is more efficient
- Can be used in a variety of complex reconstructive surgeries, including those involving radiated tissue, where graft fixation and suturing is challenging
- Future applications in endoscopic and laparoscopic surgery are possible
.jpeg)
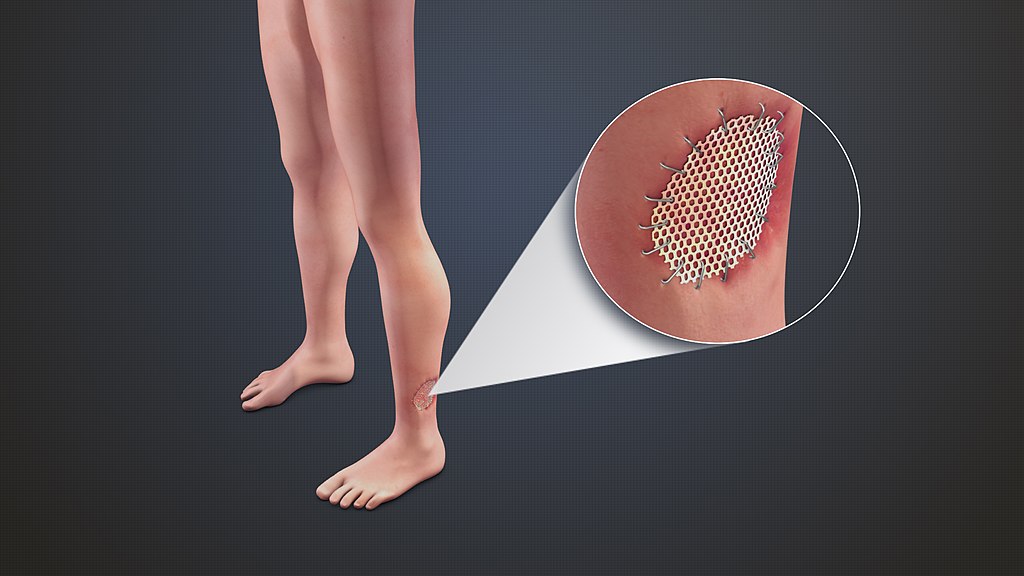
Fixation device to secure bone fragment of tibial tuberosity to native bone after osteotomy surgical...
Fixation device to secure bone fragment of tibial tuberosity to native bone after osteotomy surgical procedure. Background:
Standard osteotomy techniques to join the tibial tubercle fragment to native bone include screw fixation alone or fixation with wire or suture, which are not reliable methods to holding the bone in place to avoid displacement post-operatively and possibly leading to non-union, malunion, extensor weakness, extensor lag, or complete loss of active knee extension.Technology Overview:
Orthopedic oncology and joint reconstruction expert at Upstate Medical University has designed a device that secures the bone fragment of tibial tuberosity to the native bone using custom-made plates, screws and suture/wires after osteotomy and mobilization of the tuberosity and associated patellar tendon. https://suny.technologypublisher.com/files/sites/adobestock_322821442_(002).jpeg Advantages:
• Improves fixation of the tibial tubercle fragment by improving bone to bone healing and normal restoration of the knee.
• Reduces rate of revision surgery.
• Minimizes surgery time.
Intellectual Property Summary:
Patent Pending US 18/236,678Stage of Development:
TRL 3 - Experimental proof of concept Licensing Status:
This technology is available for licensing.




